Genome sequencing is rewriting the history of disease outbreaks – but without social context, it can
From the Black Death to the COVID-19 pandemic, combining the genetic data of a pathogen with historical records, archaeological artifacts and epidemiological studies can uncover its origins.

Fingerprinting transformed police investigations by making it possible to place a suspect at a crime scene with physical evidence. Similarly, genome sequencing has changed how disease detectives study outbreaks by allowing them to read a pathogen’s genes as a biological record of where it came from and how it spread.
One way to think about sequencing is to imagine a virus or bacteria’s genome as a recipe book. Each gene is a recipe for making a protein. When scientists sequence a pathogen, they read the order of the genetic letters in those recipes.
Over time, small changes appear in the recipes as the pathogen mutates. By comparing those changes in samples collected from different places and times, researchers can determine which infections are related and estimate when and where the pathogen entered a population.
Scientists have used sequencing in this way to track outbreaks of COVID-19, Ebola, mpox and foodborne illnesses. This information helps public health investigators connect cases that might otherwise seem unrelated.
Still, genomic sequencing has limits. It can show that different pathogen strains are related, but it cannot fully explain why an outbreak began in one place, why it spread in a particular direction, or how human behavior shaped its course. Answering those questions requires combining genomic data with historical records, archaeological artifacts, trade records and epidemiological investigations.
I am a chemist and the author of “Diseases Without Borders: Plagues, Pandemics, and Beyond,” a book for young adults on infectious disease and the ways it has shaped human history. In my research, I’ve found that while the genome can help researchers trace the evolutionary trail of a pathogen, other fields are needed to explain the environmental conditions that allowed this trail to become an outbreak.
Ancient DNA tells only part of the story
Advances in DNA sequencing and extraction over the past decade have made it possible to recover fragments of ancient DNA from bones and teeth. Researchers can use these genomes to study a metaphorical molecular fossil record of microbial evolution.
The Black Death, one of the deadliest pandemics in history, shows both the power and the limits of sequencing.
The infectious disease behind the Black Death, plague, is caused by the bacterium Yersinia pestis. DNA recovered from the teeth of people buried more than 5,000 years ago in what is now Sweden revealed the existence of an ancestral form of Y. pestis that had not yet adapted to fleas.
About 2,000 years later, the bacterium made an important evolutionary shift: It gained the ability to survive in fleas and pass back and forth between humans, rats and other mammals via flea bites. That change made the pathogen far more dangerous and helped pave the way for three great plague pandemics that followed: the Justinianic Plague from the sixth to eighth century; the Black Death and later waves from the 1300s into the 1700s; and the third pandemic from the 19th to mid-20th centuries.
But how and why did plague emerge and move through human societies with such devastating results? Genetic results alone are not enough to answer these questions.
When gravestones become genetic evidence
Geneticists needed archaeologists, paleoclimatologists and historians to complete the picture of the plague pandemics. The genome revealed the lineage. Other disciplines supplied the historical and environmental context.
Two 14th-century graveyards in what is now Kyrgyzstan provide a striking example of how historical evidence can guide genetic investigations into the origins of a pandemic.
Historian Philip Slavin noticed archival records pointing to an unusual number of gravestones from 1338 and 1339. Some of those tombstones explicitly referred to a pestilence as the cause of death.
That clue led to the next stage of the investigation, where archaeologist Maria Spyrou and her team extracted and sequenced ancient DNA from the skeletal remains of seven people buried in the graves and found genetic traces of Yersinia pestis in three of the skeletons. These strains were close precursors of the strain linked to the Black Death and ancestors of several modern Y. pestis lineages.
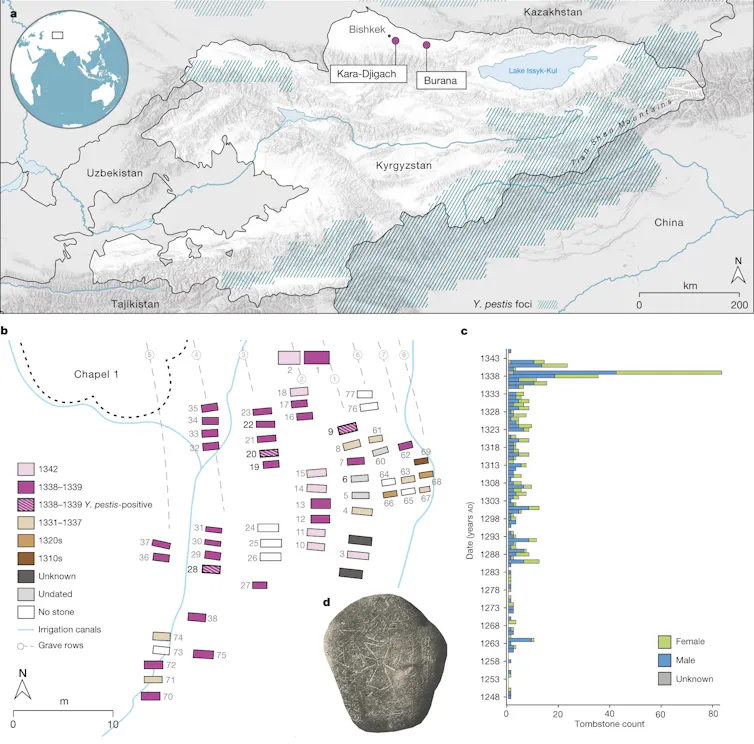
This major finding was still not the whole story. It could explain where the Black Death pandemic began but not how the disease spread across Asia to Europe. Researchers found a potential answer to this question in artifacts buried at the site, which included pearls from the Indian Ocean, Mediterranean coral and foreign coins. Those objects suggested that the region was connected to long-distance trade networks.
Once the gravestones, skeletal remains, written records and trade goods were considered together, a richer picture emerged. Researchers could place the pathogen in a specific time and place and connect it to the networks of human movement that may have carried plague westward.
Sequencing provided the biological clue, revealing the pathogen’s identity and ancestry. History and archaeology turned that clue into a plausible narrative.
From ancient DNA to modern outbreaks
Genomic sequencing isn’t limited to examining outbreak cold cases. It is also researchers’ tool of choice for understanding new diseases.
When the first reported COVID-19 cases emerged in 2019, researchers quickly sequenced the virus and found that it was closely related to the virus that caused the 2002 SARS outbreak. This placed the new virus within a known family of pathogens.
Later genomic sequencing helped reveal the scale of a major superspreading event: the 2020 Biogen conference in Boston.
The biotech company Biogen brought together about 175 European and American executives at a moment when COVID-19 was only beginning to spread in the United States. In Europe, COVID-19 was also escalating, with northern Italy reporting locally transmitted clusters just days before the meeting. After the meeting, many Massachusetts cases were linked to the conference.
Researchers then analyzed thousands of viral genomes from patients in Massachusetts and elsewhere. One viral genome carried a unique genetic signature traceable to a European attendee at the conference. It matched viruses circulating in Europe but also had an additional mutation that appeared to have arisen during the attendee’s travel to Boston or early in the conference.
Because that altered sequence appeared only in people with direct or indirect ties to the meeting, it served as a genetic marker for the COVID-19 strain originating at the Biogen conference. By comparing it with other viral sequences in national databases, researchers tracked the strain associated with the conference to 29 states and several other countries.
Interviews and contact tracing alone couldn’t have made that chain of infection so clear because people may not know exactly when they were exposed, especially when infections spread through brief encounters, via travel or large meetings.
When genomes join the investigation
Genome sequencing has rewritten the history of disease by giving scientists a way to read a pathogen’s own record of change.
It can link ancient graves to later pandemics and trace a modern outbreak from one conference room to cases across a continent.
But the greatest strength of genome sequencing lies in partnership. Sequencing does not replace history, archaeology or public health investigation. It gives them a new molecular partner.
Combining work from these fields produces a fuller and more accurate account of how disease moves through the world.
Marc Zimmer does not work for, consult, own shares in or receive funding from any company or organization that would benefit from this article, and has disclosed no relevant affiliations beyond their academic appointment.
Read These Next
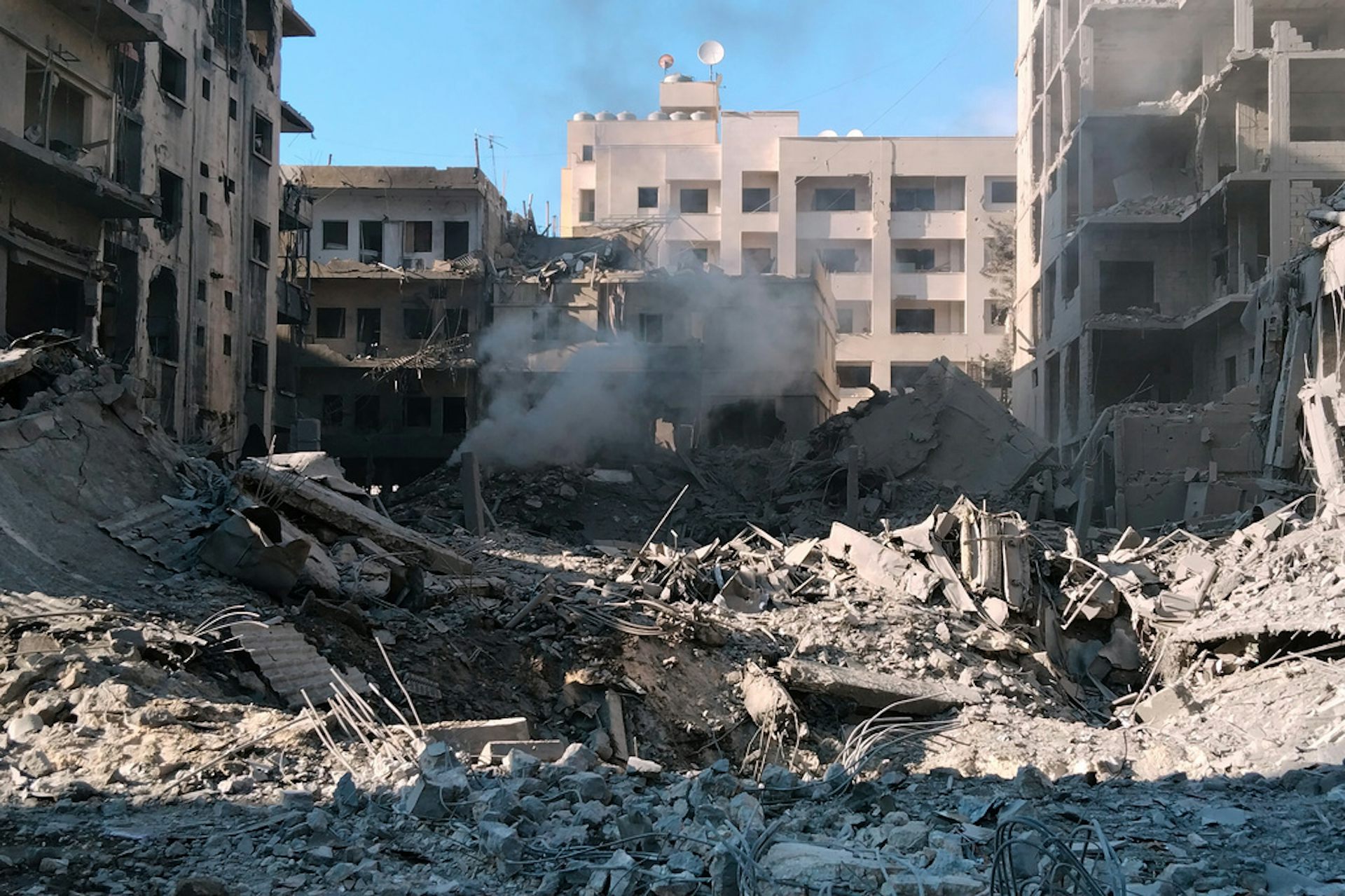
In Iran war’s shadow, Israel’s renewed Lebanon campaign risks repeating failed lessons – and occupat
Unable to defeat Iran, Israel shifts its focus to Lebanon, fearing U.S. negotiations with Tehran could…
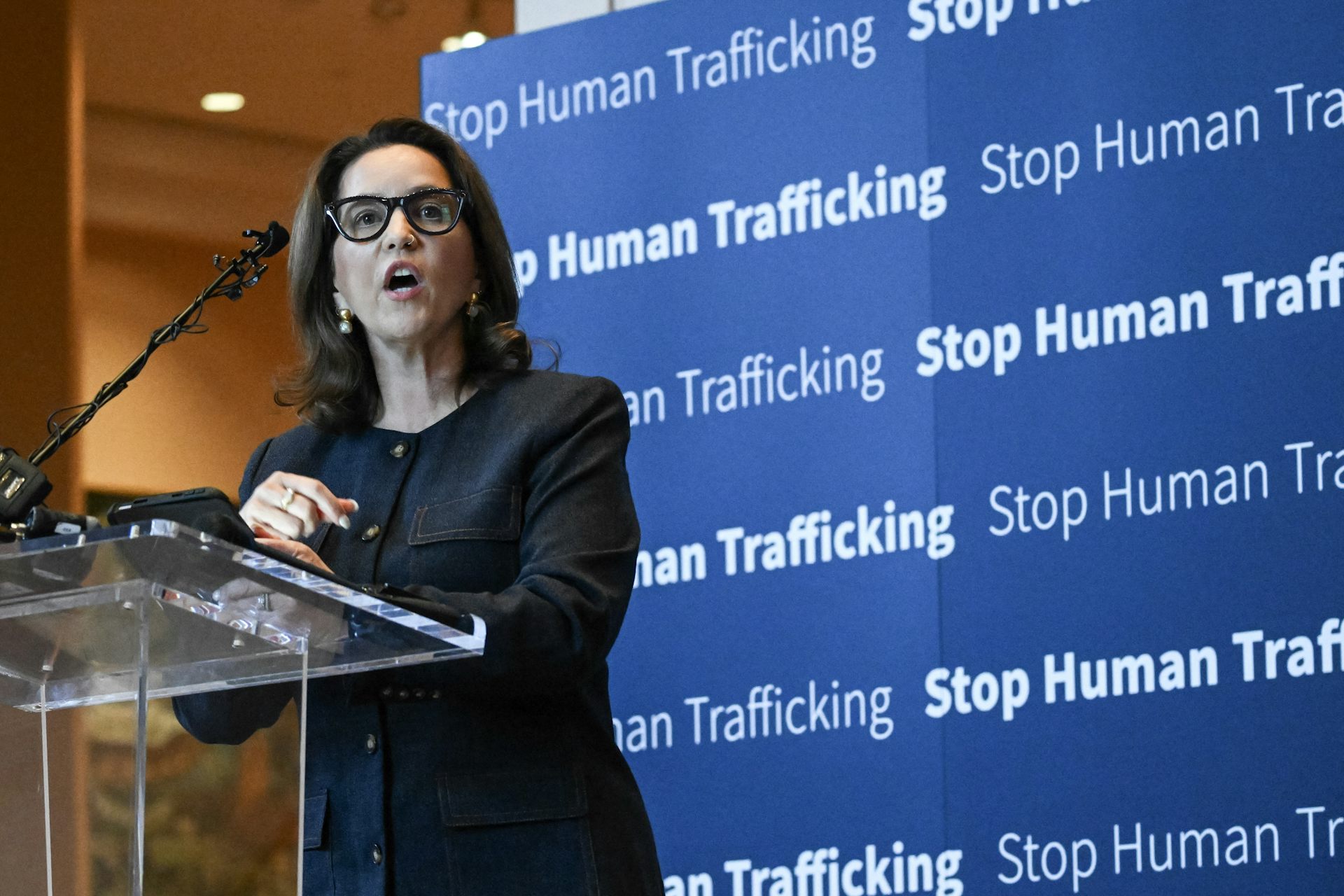
The World Cup and human trafficking: What the research reveals about the real risks at major sportin
Public awareness campaigns around the World Cup and other sporting events are well intentioned – but…
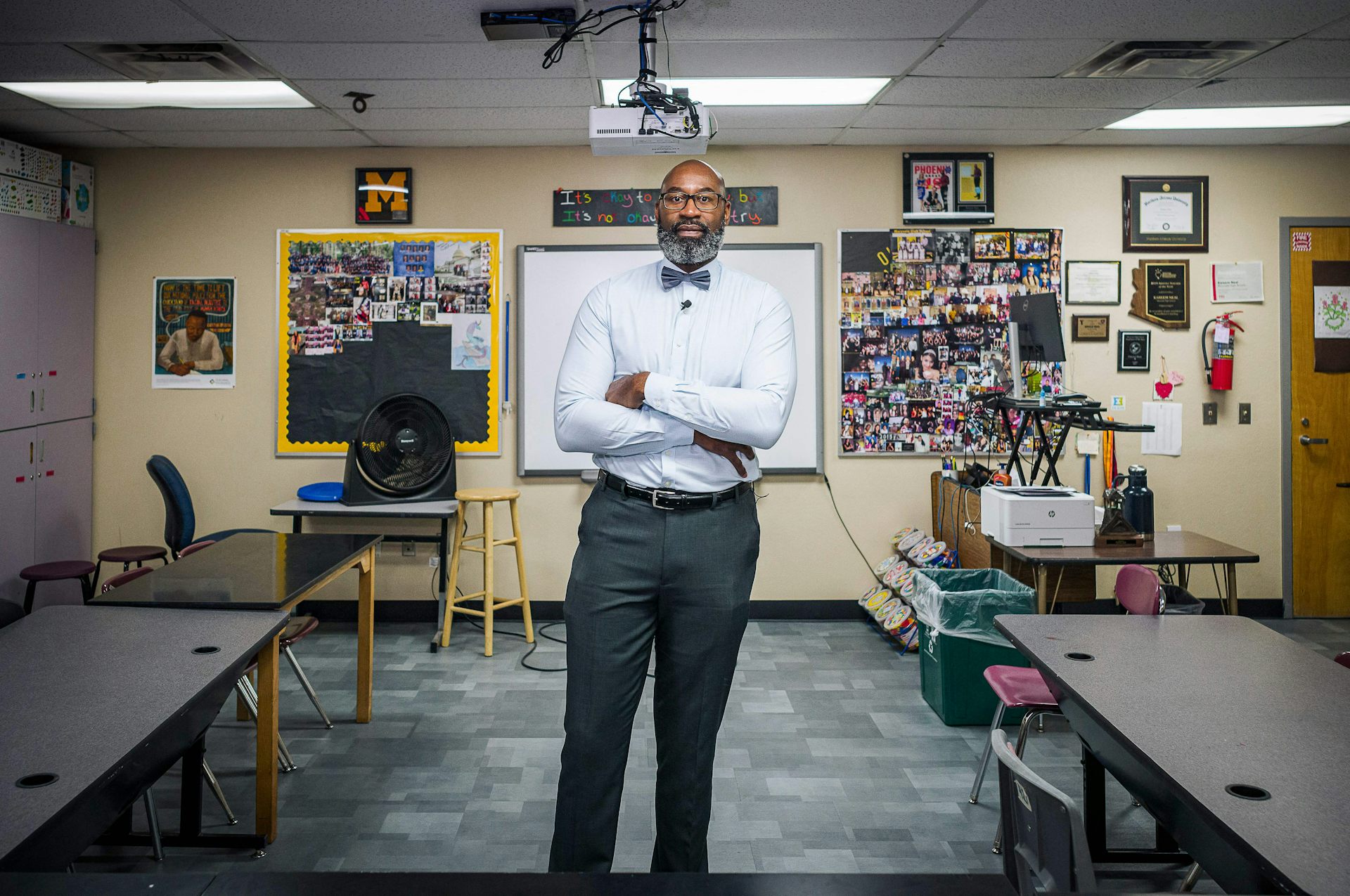
Black teachers improve outcomes for all students, but the profession remains largely white
Many Black teachers were pushed out of classrooms from the 1950s through ‘70s. Despite new recruitment…






